Neuroepigenomics of brain aging and disease
How do genetic variants for brain disorders that reside outside of genes contribute to disease?
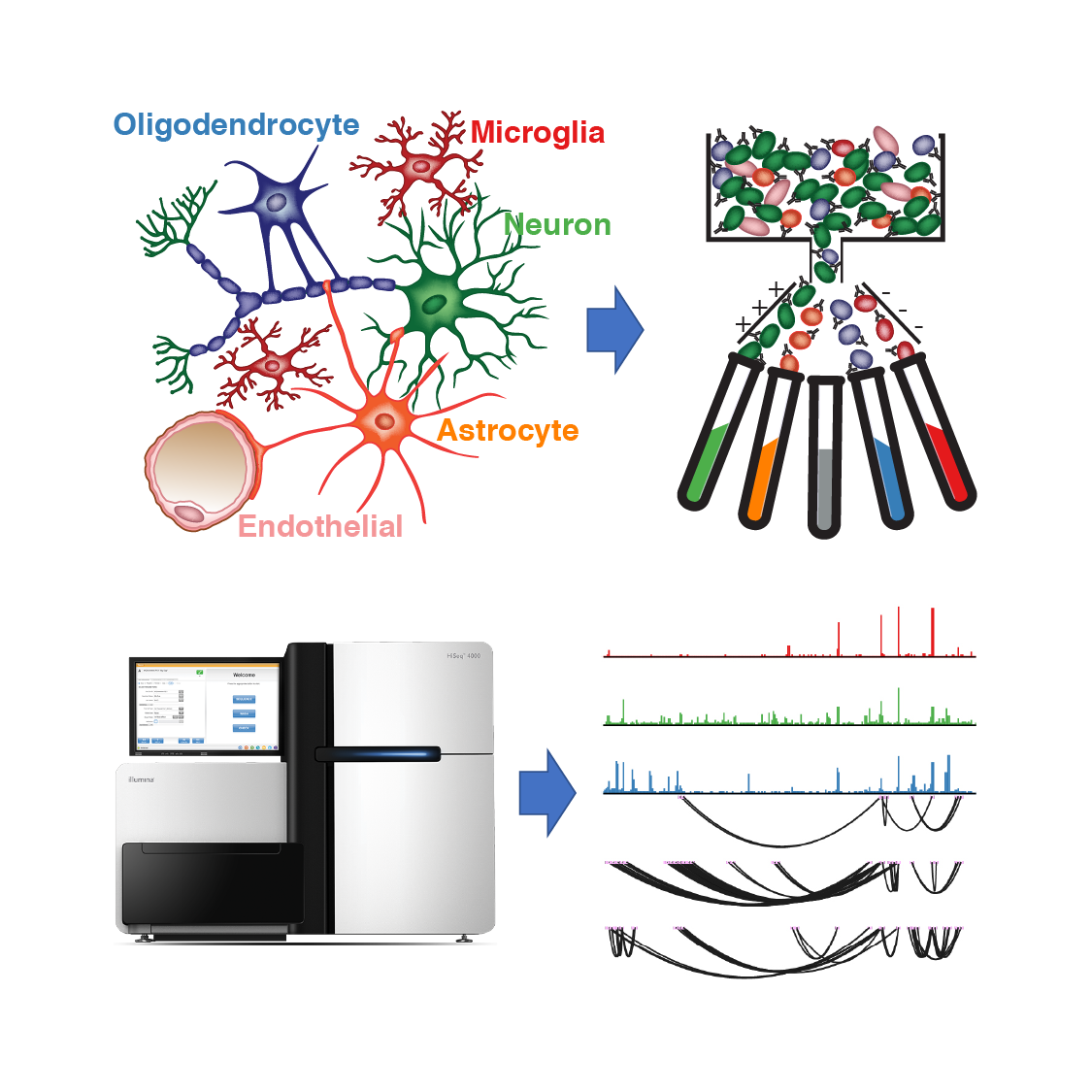
Genetic variation located outside of genes has been associated with an increased risk of aging-related brain disorders such as Alzheimer’s disease. These genetic ‘risk’ variants are thought to regulate gene expression from afar through altering the function of gene regulatory regions called enhancers. However, it is unclear which genes are affected by these variants and which brain cell types are impacted. To address this problem, we established a nuclei isolation protocol for specific cell types of the human brain.
We utilized this approach to generate enhancer-promoter interactome maps for multiple human brain cell types. We found that genetic risk variants for Alzheimer’s disease were enriched in enhancers of brain immune cells and identified their probable target genes. Lastly, we validated a microglia-specific enhancer that harbors Alzheimer’s disease-risk variants at the BIN1 locus using pluripotent stem cells and CRISPR-mediated DNA editing (Nott*, Holtman*, Coufal*, et al., Science 2019). These findings have expanded the gene repertoire influenced by Alzheimer’s disease-risk variants and revealed the probable cell types in which they function.
Degenerative diseases like Alzheimer’s disease are driven predominantly by aging. It is expected that aging associated changes of brain cell types will be driven by altered activity of enhancers. Our group will investigate how the epigenome regulates cell type gene expression programs in aging-related brain disorders using next-generation sequencing of patient-derived brain tissue and stem cell models of disease. Cell type specific epigenomic roadmaps associated with aging and disease will be utilized to identify signaling pathways and transcription factors that are dysregulated in brain disorders.
Chromatin regulators in brain disorders
How do chromatin regulators drive cell type responses to brain perturbations and contribute to disease?
Chromatin regulators modulate gene expression through altering chromatin architecture and are fundamental for conveying cellular responses to changes in the brain microenvironment. We have shown that chromatin regulators can be critical for orchestrating brain development. Specifically, histone deacetylases 2 (HDAC2), an epigenetic eraser, is negatively regulated by nitric oxide signaling and is required for normal cortical development (Nott et al. Nature 2008; Nott, et al. PNAS 2013).
We subsequently demonstrated that dysregulation of chromatin regulators can contribute to a neurodevelopmental disorder called Rett Syndrome, a disorder caused by mutations in MeCP2 (Nott et al. Nat Neurosci. 2016). We found that HDAC3, a binding partner of MeCP2, elicits Rett syndrome-associated behaviors in young mice and can activate gene expression through regulation of the transcription factor, FOXO. These molecular findings were translated to human biology using induced-pluripotent stem cells that were derived from skin cells of a Rett patient in combination with CRISPR-mediated DNA editing.
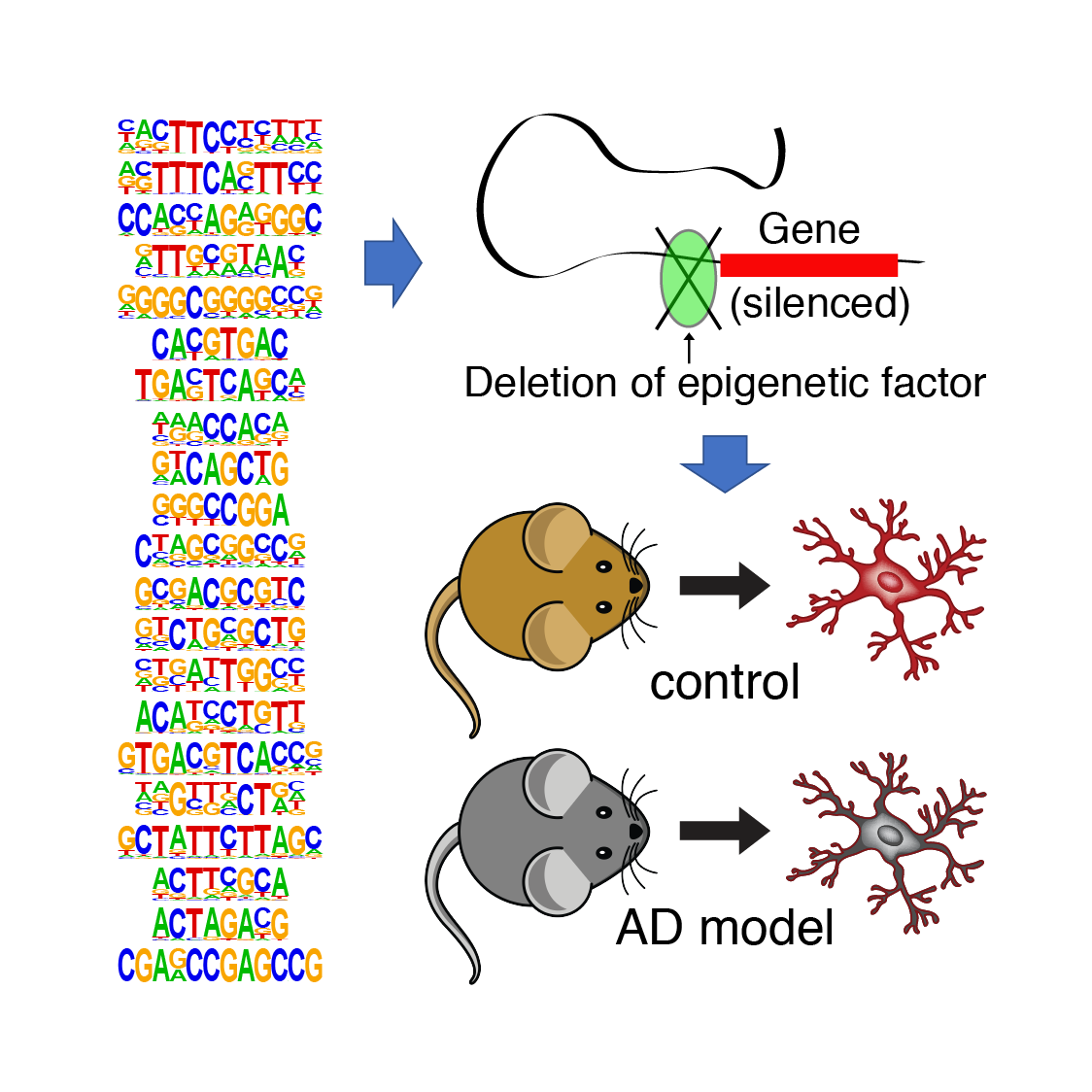
For aging-associated brain disorders, such as Alzheimer’s disease, a hallmark of pathology is neuroinflammation. Our cell type-specific epigenomic atlases identified DNA binding motifs for microglia transcription factors that are likely to be disrupted by Alzheimer’s disease-risk variants. However, the function of these microglia chromatin factors during aging-associated disease states is largely unexplored. Our group will examine the molecular and behavioral role of these putative disease-associated chromatin regulators in microglia using both mouse genetics and human stem cell-derived model systems.
Environmental impact on brain function
How do lifestyle and environment factors impact brain homeostasis and disease progression?
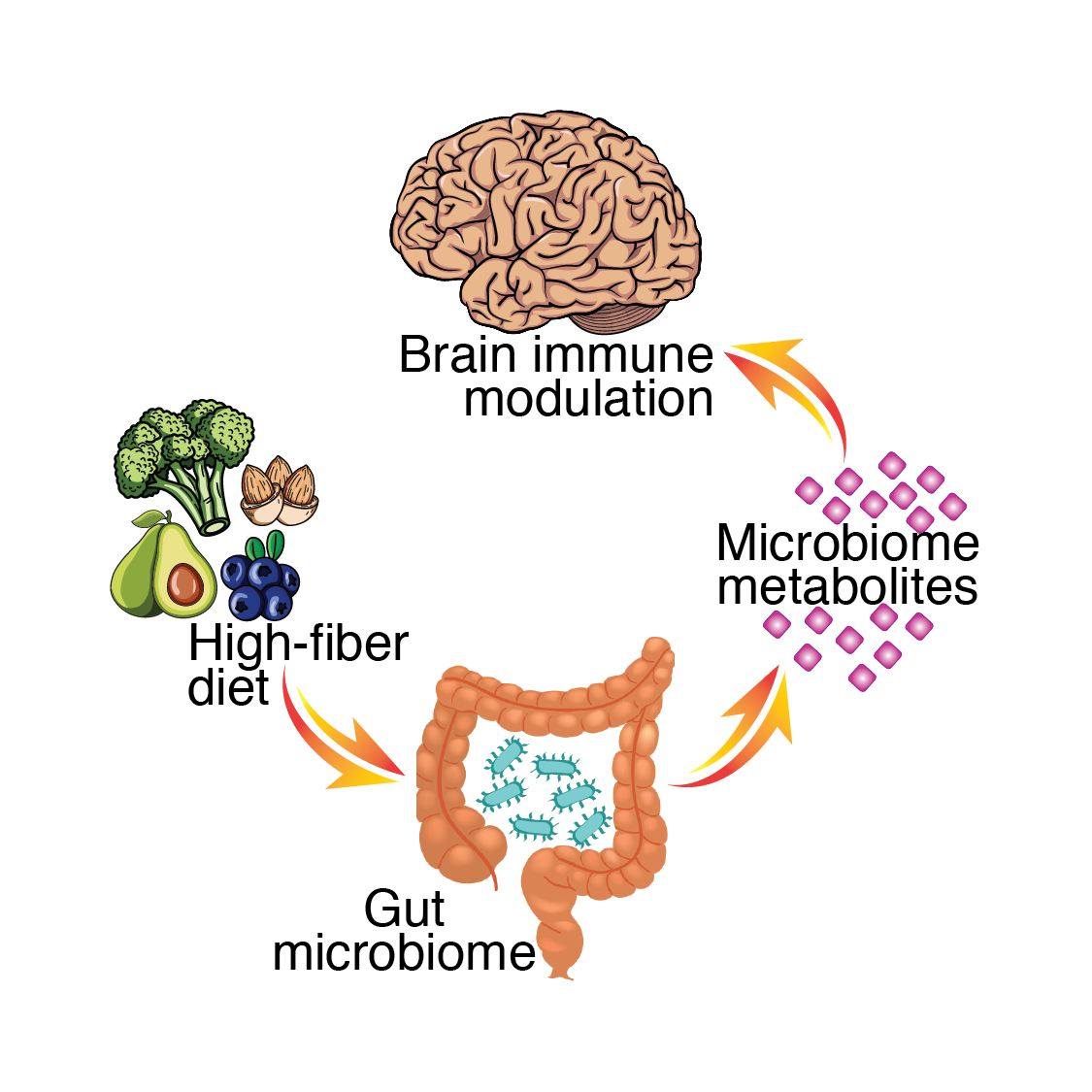
The onset, progression and severity of aging-related brain disorders can be influenced by lifetyle and environmental factors. These modifiable risk factors include diabetes, obesity, smoking, cognitive inactivity, exercise, infections and the gut microbiome. Environmental stimuli can modify cell behavior and elicit long-lasting changes that are ‘stored’ at the epigenetic level. We have shown that the epigenetic eraser, HDAC2, is required for cortical plasticity during early postnatal development when the brain is especially sensitive to environmental stimuli (Nott*, Cho*, et al., Neuroepigenetics 2015).
Aging has been associated with chronic low-grade peripheral inflammation that will impact neuroimmune function and brain health. Our group will examine the effect of environmental and lifestyle factors on the epigenome of microglia and how this impacts neuroimmune function during homeostasis, aging and disease.